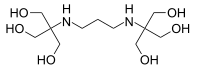
Bis-tris propane
 | |
 | |
| Names | |
|---|---|
| Other names
2,2'-(Propane-1,3-diyldiimino)bis[2-(hydroxymethyl)propane-1,3-diol] | |
| Identifiers | |
| 64431-96-5 | |
| 3D model (Jmol) | Interactive image |
| 1786109 | |
| ChEBI | CHEBI:40947 |
| ChEMBL | ChEMBL63859 |
| ChemSpider | 111383 |
| DrugBank | DB02676 |
| ECHA InfoCard | 100.058.981 |
| EC Number | 264-899-3 |
| 1734507 | |
| MeSH | 1,3-bis(tris(hydroxymethyl)methylamino)propane |
| PubChem | 125132 |
| |
| |
| Properties | |
| C11H26N2O6 | |
| Molar mass | 282.34 g·mol−1 |
| Appearance | White crystals |
| Melting point | 164 to 165 °C (327 to 329 °F; 437 to 438 K) |
| log P | −2.794 |
| Related compounds | |
| Related compounds |
|
| Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa). | |
| | |
| Infobox references | |
Bis-tris propane, or 1,3-bis(tris(hydroxymethyl)methylamino)propane, is a chemical substance that is used in buffer solutions. It is a white to off-white crystalline powder that is soluble in water. It has a wide buffering range, from 6 to 9.5 due to its two pKa values which are close in value. This buffer is primarily used in biochemistry and molecular biology.
Applications
A review of DNA polymerase fidelity cites bis-tris propane as a suitable buffer for polymerase chain reaction (PCR).[1] Bis-Tris propane has also been used with HCl buffer for stabilization of farnesyl diphosphate isolated from a strain of Saccharomyces cerevisiae.[2] It has also been used in a study of the effects of buffer identity on electric signals of light-excited bacteriorhodospin.[3] Use of Bis-Tris propane has also been documented in an investigation of the MgATPase activity of the myosin subfragment 1 monomer.[4] The effect of buffer identity on the kinetics of the restriction enzyme EcoRV has been studied in various buffers, including Bis-Tris propane.[5] Bis-Tris propane wide buffering range is also useful for calibration of genetically encoded pH indicators expressed in the cytosol or mitochondria.[6]
See also
References
- ↑ Eckert K.A. & Kunkel, T.A. (1991). "DNA polymerase fidelity and the polymerase chain reaction". PCR Methods Appl. 1 (1): 17–24. PMID 1842916.
- ↑ Song, L. (2003). "Detection of farnesyl diphosphate accumulation in yeast ERG9 mutants". Anal. Biochem. 317 (2): 180–185. doi:10.1016/S0003-2697(03)00138-6. PMID 12758256.
- ↑ Toth-Boconadi, R.; et al. (2000). "Buffer effects on electric signals of light-excited bacteriorhodospin". Biophys. J. 78 (6): 3170–3177. Bibcode:2000BpJ....78.3170T. doi:10.1016/S0006-3495(00)76853-6.
- ↑ Bachouchi N, Garrigos M, Morel JE (1986). "MgATPase activity of myosin subfragment 1. The dimer is more active than the monomer". J. Mol. Biol. 191 (2): 247–254. doi:10.1016/0022-2836(86)90261-5. PMID 2949083.
- ↑ Wenner, J.R. & Bloomfield, V.A. (1999). "Buffer effects on EcoRV kinetics as measured by fluorescent staining and digital imaging of plasmid cleavage". Anal. Biochem. 268 (2): 201–212. doi:10.1006/abio.1998.3079. PMID 10075809.
- ↑ Ivannikov, M.; et al. (2013). "Mitochondrial Free Ca2+ Levels and Their Effects on Energy Metabolism in Drosophila Motor Nerve Terminals". Biophys. J. 104 (11): 2353–2361. Bibcode:2013BpJ...104.2353I. doi:10.1016/j.bpj.2013.03.064. PMC 3672877
 . PMID 23746507.
. PMID 23746507.