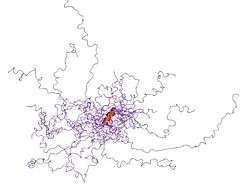
Intrinsically disordered proteins
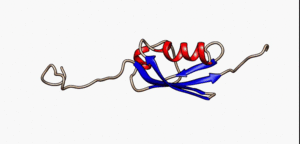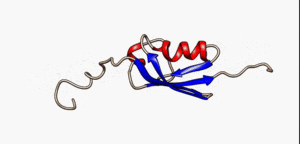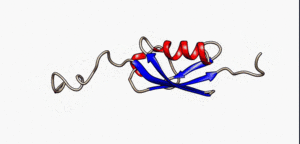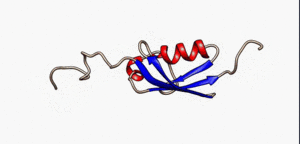
An intrinsically disordered protein (IDP) is a protein that lacks a fixed or ordered three-dimensional structure.[2][3][4] IDPs cover a spectrum of states from fully unstructured to partially structured and include random coils, (pre-)molten globules, and large multi-domain proteins connected by flexible linkers. They constitute one of the main types of protein (alongside globular, fibrous and membrane proteins).[5]
The discovery of IDPs has challenged the traditional protein structure paradigm, that protein function depends on a fixed three-dimensional structure. This dogma has been challenged over the last decades by increasing evidence from various branches of structural biology, suggesting that protein dynamics may be highly relevant for such systems. Despite their lack of stable structure, IDPs are a very large and functionally important class of proteins. In some cases, IDPs can adopt a fixed three-dimensional structure after binding to other macromolecules. Overall, IDPs are different from structured proteins in many ways and tend to have distinct properties in terms of function, structure, sequence, interactions, evolution and regulation.[6]
History
In the 1930s -1950s, the first protein structures were solved by protein crystallography. These early structures suggested that a fixed three-dimensional structure might be generally required to mediate biological functions of proteins. When stating that proteins have just one uniquely defined configuration, Mirsky and Pauling did not recognize that Fisher's work would have supported their thesis with his 'Lock and Key' model (1894). These publications solidified the central dogma of molecular biology in that the sequence determines the structure which, in turn, determines the function of proteins. In 1950, Karush wrote about 'Configurational Adaptability' contradicting all the assumptions and research in the 19th century. He was convinced that proteins have more than one configuration at the same energy level and can choose one when binding to other substrates. In the 1960s, Levinthal's paradox suggested that the systematic conformational search of a long polypeptide is unlikely to yield a single folded protein structure on biologically relevant timescales (i.e. seconds to minutes). Curiously, for many (small) proteins or protein domains, relatively rapid and efficient refolding can be observed in vitro. As stated in Anfinsen's Dogma from 1973, the fixed 3D structure of these proteins is uniquely encoded in its primary structure (the amino acid sequence), is kinetically accessible and stable under a range of (near) physiological conditions, and can therefore be considered as the native state of such "ordered" proteins.
During the subsequent decades, however, many large protein regions could not be assigned in x-ray datasets, indicating that they occupy multiple positions which average out in electron density maps. The lack of a fixed, unique positions relative to the crystal lattice suggested that these regions were "disordered". Additional techniques for determining protein structures, such as NMR, demonstrated the presence of large flexible linkers and termini in many solved structural ensembles. It is now generally accepted that proteins exist as an ensemble of similar structures with some regions more constrained than others. Intrinsically Unstructured Proteins (IUPs) occupy the extreme end of this spectrum of flexibility, whereas IDPs also include proteins of considerable local structure tendency or flexible multidomain assemblies.These highly dynamic disordered regions of proteins have subsequently been linked to functionally important phenomena such as allosteric regulation and enzyme catalysis. [8] [9]
In the 2000s, bioinformatic predictions of intrinsic disorder in proteins indicated that intrinsic disorder is more common in sequenced/predicted proteomes than in known structures in the protein database. Based on DISOPRED2 prediction, long (>30 residue) disordered segments occur in 2.0% of archaean, 4.2% of eubacterial and 33.0% of eukaryotic proteins.[10] In 2001, Dunker published his paper 'Intrinsically Disordered Proteins' questioning whether the newly found information was ignored for 50 years.[11]
In the 2010s it became clear that IDPs are highly abundant among disease-related proteins.[12]
Biological roles
Many disordered proteins have the binding affinity with their receptors regulated by post-translational modification, thus it has been proposed that the flexibility of disordered proteins facilitates the different conformational requirements for binding the modifying enzymes as well as their receptors.[13] Intrinsic disorder is particularly enriched in proteins implicated in cell signaling, transcription and chromatin remodeling functions.[14][15]
Flexible linkers
Disordered regions are often found as flexible linkers or loops connecting domains. Linker sequences vary greatly in length but are typically rich in polar uncharged amino acids . Flexible linkers allow the connecting domains to freely twist and rotate to recruit their binding partners via protein domain dynamics. They also allow their binding partners to induce larger scale conformational changes by long-range allostery
Linear motifs
Linear motifs are short disordered segments of proteins that mediate functional interactions with other proteins or other biomolecules (RNA, DNA, sugars etc.). Many roles or linear motifs are associated with cell regulation, for instance in control of cell shape, subcellular localisation of individual proteins and regulated protein turnover. Often, post-translational modifications such as phosphorylation tune the affinity (not rarely by several orders of magnitude) of individual linear motifs for specific interactions. Relatively rapid evolution and a relatively small number of structural restraints for establishing novel (low-affinity) interfaces make it particularly challenging to detect linear motifs but their widespread biological roles and the fact that many viruses mimick/hijack linear motifs to efficiently recode infected cells underlines the timely urgency of research on this very challenging and exciting topic. Unlike globular proteins IDPs do not have spatially-disposed active pockets. Nevertheless, in 80% of IDPs (~3 dozens) subjected to detailed structural characterization by NMR there are linear motifs termed PreSMos (pre-structured motifs) that are transient secondary structural elements primed for target recognition. In several cases it has been demonstrated that these transient structures become full and stable secondary structures, e.g., helices, upon target binding. Hence, PreSMos are the putative active sites in IDPs.[16]
Coupled folding and binding
Many unstructured proteins undergo transitions to more ordered states upon binding to their targets (e.g. Molecular Recognition Features (MoRFs)[17]). The coupled folding and binding may be local, involving only a few interacting residues, or it might involve an entire protein domain. It was recently shown that the coupled folding and binding allows the burial of a large surface area that would be possible only for fully structured proteins if they were much larger.[18] Moreover, certain disordered regions might serve as "molecular switches" in regulating certain biological function by switching to ordered conformation upon molecular recognition like small molecule-binding, DNA/RNA binding, ion interactions etc.[19]
The ability of disordered proteins to bind, and thus to exert a function, shows that stability is not a required condition. Many short functional sites, for example Short Linear Motifs are over-represented in disordered proteins.
Disorder in the bound state (fuzzy complexes)
Intrinsically disordered proteins can retain their conformational freedom even when they bind specifically to other proteins. The structural disorder in bound state can be static or dynamic. In fuzzy complexes structural multiplicity is required for function and the manipulation of the bound disordered region changes activity. The conformational ensemble of the complex is modulated via post-translational modifications or protein interactions.[20] Specificity of DNA binding proteins often depends on the length of fuzzy regions, which is varied by alternative splicing.[21]
Structural Aspects
Intrinsically disordered proteins adapt many different structures in vivo according to the cell's conditions, creating a structural or conformational ensemble.[22][23]
Therefore their structures are strongly function-related. However, only few proteins are fully disordered in their native state. Disorder is mostly found in intrinsically disordered regions (IDRs) within an otherwise well-structured protein. The term intrinsically disordered protein (IDP) therefore includes proteins that contain IDRs as well as fully disordered proteins.
The existence and kind of protein disorder is encoded in its amino acid sequence.[2] In general, IDPs are characterized by a low content of bulky hydrophobic amino acids and a high proportion of polar and charged amino acids, usually referred to as low hydrophobicity.[22] This property leads to good interactions with water. Furthermore high net charges promote disorder because of electrostatic repulsion resulting from equally charged residues.[23] Thus disordered sequences cannot sufficiently bury a hydrophobic core to fold into stable globular proteins. In some cases, hydrophobic clusters in disordered sequences provide the clues for identifying the regions that undergo coupled folding and binding (refer to biological roles). Many disordered proteins reveal regions without any regular secondary structure These regions can be termed as flexible, compared to structured loops. While the latter are rigid and contain only one set of Ramachandran angles, IDPs involve multiple sets of angles.[23] The term flexibility is also used for well-structured proteins, but describes a different phenomenon in the context of disordered proteins. Flexibility in structured proteins is bound to an equilibrium state, while it is not so in IDPs.[23] Many disordered proteins also reveal low complexity sequences, i.e. sequences with over-representation of a few residues. While low complexity sequences are a strong indication of disorder, the reverse is not necessarily true, that is, not all disordered proteins have low complexity sequences. Disordered proteins have a low content of predicted secondary structure.
Experimental validation
Intrinsically unfolded proteins, once purified, can be identified by various experimental methods. The primary method to obtain information on disordered regions of a protein is NMR spectroscopy. The lack of electron density in X-ray crystallographic studies may also be a sign of disorder.
Folded proteins have a high density (partial specific volume of 0.72-0.74 mL/g) and commensurately small radius of gyration. Hence, unfolded proteins can be detected by methods that are sensitive to molecular size, density or hydrodynamic drag, such as size exclusion chromatography, analytical ultracentrifugation, Small angle X-ray scattering (SAXS), and measurements of the diffusion constant. Unfolded proteins are also characterized by their lack of secondary structure, as assessed by far-UV (170-250 nm) circular dichroism (esp. a pronounced minimum at ~200 nm) or infrared spectroscopy. Unfolded proteins also have exposed backbone peptide groups exposed to solvent, so that they are readily cleaved by proteases, undergo rapid hydrogen-deuterium exchange and exhibit a small dispersion (<1 ppm) in their 1H amide chemical shifts as measured by NMR. (Folded proteins typically show dispersions as large as 5 ppm for the amide protons.) Recently, new methods including Fast parallel proteolysis (FASTpp) have been introduced, which allow to determine the fraction folded/disordered without the need for purification.[24][25] Even subtle differences in the stability of missense mutations, protein partner binding and (self)polymerisation-induced folding of (e.g.) coiled-coils can be detected using FASTpp as recently demonstrated using the tropomyosin-troponin protein interaction.[26] Fully unstructured protein regions can be experimentally validated by their hypersusceptibility to proteolysis using short digestion times and low protease concentrations.[27]
Bulk methods to study IDP structure and dynamics include SAXS for ensemble shape information, NMR for atomistic ensemble refinement, Fluorescence for visualising molecular interactions and conformational transitions, x-ray crystallography to highlight more mobile regions in otherwise rigid protein crystals, cryo-EM to reveal less fixed parts of proteins, light scattering to monitor size distributions of IDPs or their aggregation kinetics, Circular Dichroism to monitor secondary structure of IDPs.
Single-molecule methods to study IDPs include spFRET[28] to study conformational flexibility of IDPs and the kinetics of structural transitions, optical tweezers[29] for high-resolution insights into the ensembles of IDPs and their oligomers or aggregates, nanopores[30] to reveal global shape distributions of IDPs, magnetic tweezers[31] to study structural transitions for long times at low forces, high-speed AFM[32] to visualise the spatio-temporal flexibility of IDPs directly.
Disorder prediction
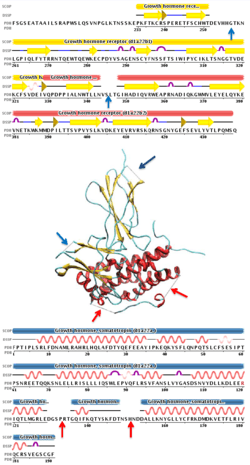
Disorder prediction algorithms can predict Intrinsic Disorder (ID) propensity with high accuracy (approaching around 80%) based on primary sequence composition, similarity to unassigned segments in protein x-ray datasets, flexible regions in NMR studies and physico-chemical properties of amino acids.
Distinguishing IDPs from well-structured proteins
Separating disordered from ordered proteins is essential for disorder prediction. One of the first steps to find a factor that distinguishes IDPs from non-IDPs is to specify biases within the amino acid composition. The following hydrophilic, charged amino acids A, R, G, Q, S, P, E and K have been characterized as disorder-promoting amino acids, while order-promoting amino acids W, C, F, I, Y, V, L, and N are hydrophobic and uncharged. The remaining amino acids H, M, T and D are ambiguous, found in both ordered and unstructured regions.[2] This information is the basis of most sequence-based predictors. Regions with little to no secondary structure, also known as NORS (NO Regular Secondary structure) regions,[33] and low-complexity regions can easily be detected. However, not all disordered proteins contain such low complexity sequences.
Prediction methods
Determining disordered regions from biochemical methods is very costly and time-consuming. Due to the variable nature of IDPs, only certain aspects of their structure can be detected, so that a full characterization requires a large number of different methods and experiments. This further increases the expense of IDP determination. In order to overcome this obstacle, computer-based methods are created for predicting protein structure and function. It is one of the main goals of bioinformatics to derive knowledge by prediction. Predictors for IDP function are also being developed, but mainly use structural information such as linear motif sites.[4][34] There are different approaches for predicting IDP structure, such as neural networks or matrix calculations, based on different structural and/or biophysical properties.
Many computational methods exploit sequence information to predict whether a protein is disordered.[35] Notable examples of such software include IUPRED and Disopred. Different methods may use different definitions of disorder. Meta-predictors show a new concept, combining different primary predictors to create a more competent and exact predictor.
Due to the different approaches of predicting disordered proteins, estimating their relative accuracy is fairly difficult. For example, neural networks are often trained on different datasets. The disorder prediction category is a part of biannual CASP experiment that is designed to test methods according accuracy in finding regions with missing 3D structure (marked in PDB files as REMARK465, missing electron densities in X-ray structures).
Disorder and disease
Intrinsically unstructured proteins have been implicated in a number of diseases.[36] Aggregation of misfolded proteins is the cause of many synucleinopathies and toxicity as those proteins start binding to each other randomly and can lead to cancer or cardiovascular diseases. Thereby, misfolding can happen spontaneously because millions of copies of proteins are made during the lifetime of an organism. The aggregation of the intrinsically unstructured protein α-Synuclein is thought to be responsible. The structural flexibility of this protein together with its susceptibility to modification in the cell leads to misfolding and aggregation. Genetics, oxidative and nitrative stress as well as mitochondrial impairment impact the structural flexibility of the unstructured α-Synuclein protein and associated disease mechanisms.[37] Many key oncogenes have large intrinsically unstructured regions, for example p53 and BRCA1. These regions of the proteins are responsible for mediating many of their interactions. Taking the cell's native defense mechanisms as a model drugs can be developed, trying to block the place of noxious substrates and inhibiting them, and thus counteracting the disease.[38]
Computer simulations
Structural and dynamical properties of intrinsically unstructured proteins are being studied by molecular dynamics simulations.[39][40][41] Findings from these simulations suggest a highly flexible conformational ensemble of intrinsically disordered proteins at different temperatures which is related to the presence of low free energy barriers.
Effects of confinement have also recently been addressed.[42] For example, these studies suggest that confinement tends to increase the population of turn structures with respect to the population of coils and β-hairpins.
Moreover, various protocols and methods of analyzing IDPs, such as studies based on quantitative analysis of GC content in genes and their respective chromosomal bands, have been used to understand functional IDP segments.[43][44]
Pioneering IDP research labs
In the last ten years, a great number of laboratories have investigated protein disorders using both experimental (e.g. SAXS-NMR, single-molecule fluorescence) and computational (analysis of protein structure) techniques.
See also
References
- ↑ Majorek K, Kozlowski L, Jakalski M, Bujnicki, JM (December 18, 2008). "Chapter 2: First Steps of Protein Structure Prediction". In Bujnicki, J. Prediction of Protein Structures, Functions, and Interactions (PDF). John Wiley & Sons, Ltd. pp. 39–62. doi:10.1002/9780470741894.ch2. ISBN 9780470517673.
- 1 2 3 Dunker, A. K.; Lawson, J. D.; Brown, C. J.; Williams, R. M.; Romero, P; Oh, J. S.; Oldfield, C. J.; Campen, A. M.; Ratliff, C. M.; Hipps, K. W.; Ausio, J; Nissen, M. S.; Reeves, R; Kang, C; Kissinger, C. R.; Bailey, R. W.; Griswold, M. D.; Chiu, W; Garner, E. C.; Obradovic, Z (2001). "Intrinsically disordered protein". Journal of molecular graphics & modelling. 19 (1): 26–59. doi:10.1016/s1093-3263(00)00138-8. PMID 11381529.
- ↑ Dyson HJ, Wright PE (March 2005). "Intrinsically unstructured proteins and their functions". Nat. Rev. Mol. Cell Biol. 6 (3): 197–208. doi:10.1038/nrm1589. PMID 15738986.
- 1 2 Dunker AK, Silman I, Uversky VN, Sussman JL (December 2008). "Function and structure of inherently disordered proteins". Curr. Opin. Struct. Biol. 18 (6): 756–64. doi:10.1016/j.sbi.2008.10.002. PMID 18952168.
- ↑ Andreeva, A (2014). "SCOP2 prototype: a new approach to protein structure mining". Nucleic Acids Res. 42: D310–4. doi:10.1093/nar/gkt1242. PMC 3964979
 . PMID 24293656.
. PMID 24293656. - ↑ van der Lee, Robin; Buljan, Marija; Lang, Benjamin; Weatheritt, Robert J.; Daughdrill, Gary W.; Dunker, A. Keith; Fuxreiter, Monika; Gough, Julian; et al. (2014-07-09). "Classification of Intrinsically Disordered Regions and Proteins". Chemical Reviews. 114 (13): 6589–6631. doi:10.1021/cr400525m. ISSN 0009-2665. PMC 4095912
 . PMID 24773235.
. PMID 24773235. - ↑ Song J, Lee MS, Carlberg I, Vener AV, Markley JL (December 2006). "Micelle-induced folding of spinach thylakoid soluble phosphoprotein of 9 kDa and its functional implications". Biochemistry. 45 (51): 15633–43. doi:10.1021/bi062148m. PMC 2533273
 . PMID 17176085.
. PMID 17176085. - ↑ Bu Z, Callaway DJ (2011). "Proteins MOVE! Protein dynamics and long-range allostery in cell signaling". Advances in Protein Chemistry and Structural Biology. Advances in Protein Chemistry and Structural Biology. 83: 163–221. doi:10.1016/B978-0-12-381262-9.00005-7. ISBN 9780123812629. PMID 21570668.
- ↑ Kamerlin, S. C.; Warshel, A (2010). "At the dawn of the 21st century: Is dynamics the missing link for understanding enzyme catalysis?". Proteins: Structure, Function, and Bioinformatics. 78 (6): 1339–75. doi:10.1002/prot.22654. PMC 2841229
 . PMID 20099310.
. PMID 20099310. - ↑ Ward, J. J.; Sodhi, J. S.; McGuffin, L. J.; Buxton, B. F.; Jones, D. T. (2004). "Prediction and functional analysis of native disorder in proteins from the three kingdoms of life". Journal of Molecular Biology. 337 (3): 635–45. doi:10.1016/j.jmb.2004.02.002. PMID 15019783.
- ↑ Dunker, A. K.; Lawson, J. D.; Brown, C. J.; Williams, R. M.; Romero, P.; Oh, J. S.; Oldfield, C. J.; Campen, A. M.; Ratliff, C. M. (2001-01-01). "Intrinsically disordered protein". Journal of Molecular Graphics & Modelling. 19 (1): 26–59. doi:10.1016/s1093-3263(00)00138-8. ISSN 1093-3263. PMID 11381529.
- ↑ Uversky, V. N.; Oldfield, C. J.; Dunker, A. K. (2008). "Intrinsically Disordered Proteins in Human Diseases: Introducing the D2Concept". Annual Review of Biophysics. 37: 215–246. doi:10.1146/annurev.biophys.37.032807.125924. PMID 18573080.
- ↑ Collins MO, Yu L, Campuzano I, Grant SG, Choudhary JS (July 2008). "Phosphoproteomic analysis of the mouse brain cytosol reveals a predominance of protein phosphorylation in regions of intrinsic sequence disorder". Mol. Cell Proteomics. 7 (7): 1331–48. doi:10.1074/mcp.M700564-MCP200. PMID 18388127.
- ↑ Iakoucheva LM, Brown CJ, Lawson JD, Obradović Z, Dunker AK (October 2002). "Intrinsic disorder in cell-signaling and cancer-associated proteins". J. Mol. Biol. 323 (3): 573–84. doi:10.1016/S0022-2836(02)00969-5. PMID 12381310.
- ↑ Sandhu KS (2009). "Intrinsic disorder explains diverse nuclear roles of chromatin remodeling proteins". J. Mol. Recognit. 22 (1): 1–8. doi:10.1002/jmr.915. PMID 18802931.
- ↑ Lee, SH.; Kim, DH.; Han, JJ.; Cha, EJ.; Lim, JE.; Cho, YJ.; Lee, C.; Han, KH. (2012). "Understanding pre-structured motifs (PreSMos) in intrinsically unfolded proteins.". Curr Protein Pept Sci. 13 (1): 34–54. doi:10.2174/138920312799277974. PMID 22044148.
- ↑ Amrita Mohan; Christopher J. Oldfield; Predrag Radivojac; Vladimir Vacic; Marc S. Cortese; A. Keith Dunker; Vladimir N. Uversky. "Analysis of Molecular Recognition Features (MoRFs)". Journal of Molecular Biology 2006 Oct 6. 362(5): 1043–59. doi:10.1016/j.jmb.2006.07.087. PMID 16935303.
- ↑ Gunasekaran K, Tsai CJ, Kumar S, Zanuy D, Nussinov R (February 2003). "Extended disordered proteins: targeting function with less scaffold". Trends Biochem. Sci. 28 (2): 81–5. doi:10.1016/S0968-0004(03)00003-3. PMID 12575995.
- ↑ Sandhu KS, Dash D (July 2007). "Dynamic alpha-helices: conformations that do not conform". Proteins. 68 (1): 109–22. doi:10.1002/prot.21328. PMID 17407165.
- ↑ Fuxreiter, M (2012). "Fuzziness: Linking regulation to protein dynamics". Molecular bioSystems. 8 (1): 168–77. doi:10.1039/c1mb05234a. PMID 21927770.
- ↑ Fuxreiter, M; Simon, I; Bondos, S (2011). "Dynamic protein-DNA recognition: Beyond what can be seen". Trends in Biochemical Sciences. 36 (8): 415–23. doi:10.1016/j.tibs.2011.04.006. PMID 21620710.
- 1 2 Uversky, V. N. (2011). "Intrinsically disordered proteins from A to Z". The International Journal of Biochemistry and Cell Biology. 43: 1090–1103. doi:10.1016/j.biocel.2011.04.001.
- 1 2 3 4 Oldfield, C. (2014). "Intrinsically Disordered Proteins and Intrinsically Disordered Protein Regions". Annual Review of Biochemistry. 83: 553–584. doi:10.1146/annurev-biochem-072711.
- ↑ Minde, David P.; Maurice, Madelon M.; Rüdiger, Stefan G. D. (2012). Uversky, Vladimir N, ed. "Determining Biophysical Protein Stability in Lysates by a Fast Proteolysis Assay, FASTpp". PLoS ONE. 7 (10): e46147. doi:10.1371/journal.pone.0046147. PMC 3463568
 . PMID 23056252.
. PMID 23056252. - ↑ Park, C; Marqusee, S (2005). "Pulse proteolysis: A simple method for quantitative determination of protein stability and ligand binding". Nature Methods. 2 (3): 207–12. doi:10.1038/nmeth740. PMID 15782190.
- ↑ Robaszkiewicz, K.; Ostrowska, Z.; Cyranka-Czaja, A.; Moraczewska, J. (2015). "Impaired tropomyosin–troponin interactions reduce activation of the actin thin filament". Biochimica et Biophysica Acta (BBA) - Proteins and Proteomics. 1854: 381–390. doi:10.1016/j.bbapap.2015.01.004.
- ↑ Minde, David P.; Radli, Martina; Forneris, Frederico; Maurice, Madelon M.; Rüdiger, Stefan G. D. (2013). Buckle, Ashley M, ed. "Large Extent of Disorder in Adenomatous Polyposis Coli Offers a Strategy to Guard Wnt Signalling against Point Mutations". PLoS ONE. 8 (10): e77257. doi:10.1371/journal.pone.0077257. PMC 3793970
 . PMID 24130866.
. PMID 24130866. - ↑ Brucale, M; Schuler, B; Samorì, B (2014). "Single-Molecule Studies of Intrinsically Disordered Proteins". Chemical Reviews. 114 (6): 140117081415006. doi:10.1021/cr400297g. PMID 24432838.
- ↑ Neupane, K; Solanki, A; Sosova, I; Belov, M; Woodside, M. T. (2014). "Diverse metastable structures formed by small oligomers of α-synuclein probed by force spectroscopy". PLoS ONE. 9 (1): e86495. doi:10.1371/journal.pone.0086495. PMC 3901707
 . PMID 24475132.
. PMID 24475132. - ↑ Japrung, D; Dogan, J; Freedman, K. J.; Nadzeyka, A; Bauerdick, S; Albrecht, T; Kim, M. J.; Jemth, P; Edel, J. B. (2013). "Single-molecule studies of intrinsically disordered proteins using solid-state nanopores". Analytical Chemistry. 85 (4): 2449–56. doi:10.1021/ac3035025. PMID 23327569.
- ↑ Min, D; Kim, K; Hyeon, C; Cho, Y. H.; Shin, Y. K.; Yoon, T. Y. (2013). "Mechanical unzipping and rezipping of a single SNARE complex reveals hysteresis as a force-generating mechanism". Nature Communications. 4 (4): 1705. doi:10.1038/ncomms2692. PMC 3644077
 . PMID 23591872.
. PMID 23591872. - ↑ Miyagi, A; Tsunaka, Y; Uchihashi, T; Mayanagi, K; Hirose, S; Morikawa, K; Ando, T (2008). "Visualization of intrinsically disordered regions of proteins by high-speed atomic force microscopy". ChemPhysChem. 9 (13): 1859–66. doi:10.1002/cphc.200800210. PMID 18698566.
- ↑ Schlessinger, A.; Schaefer, C.; Vicedo, E.; Schmidberger, M.; Punta, M.; Rost, B. (2011). "Protein disorder - a breakthrough invention of evolution?". Current Opinion in Structural Biology. 21: 412–418. doi:10.1016/j.sbi.2011.03.14.
- ↑ Tompa, P. (2011). "Unstructural biology coming of age". Current Opinion in Structural Biology. 21: 419–425. doi:10.1016/j.sbi.2011.03.12.
- ↑ Ferron F, Longhi S, Canard B, Karlin D (October 2006). "A practical overview of protein disorder prediction methods". Proteins. 65 (1): 1–14. doi:10.1002/prot.21075. PMID 16856179.
- ↑ Uversky, Vladimir N.; Oldfield, Christopher J.; Dunker, A. Keith (2008). "Intrinsically Disordered Proteins in Human Diseases: Introducing the D2Concept". Annual Review of Biophysics. 37: 215–46. doi:10.1146/annurev.biophys.37.032807.125924. PMID 18573080.
- ↑ Coskuner, Orkid; Wise-Scira, Olivia; Dunn, Aquila (2013). "Structures of the E46K Mutant-Type α-Synuclein Protein and Impact of E46K Mutation on the Structures of the Wild-Type α-Synuclein Protein". ACS Chemical Neuroscience. 4: 498–508. doi:10.1021/cn3002027.
- ↑ Dobson, Christopher M. (2003-12-18). "Protein folding and misfolding". Nature. 426 (6968): 884–890. doi:10.1038/nature02261. ISSN 1476-4687. PMID 14685248.
- ↑ Mittal J, Yoo T, Georgiou G, Truskett T (December 2013). "Structural Ensemble of an Intrinsically Disordered Polypeptide". J. Of Phys. Chem B. 117: 118–124. doi:10.1021/jp308984e.
- ↑ Ojeda-May P, Pu J (August 2013). "Replica exchange molecular dynamics simulations of an α/β-type small acid soluble protein (SASP)". Biophysical Chemistry. 184: 17–21. doi:10.1016/j.bpc.2013.07.014. PMID 24029407.
- ↑ Higo J, Ito H, Kuroda M, Ono S, Nakajima N, Nakamura H (March 2001). "Energy landscape of a peptide consisting of α-helix, 3-10-helix, β-hairpin, and other disordered conformations". Protein Science. 10 (6): 1160–1171. doi:10.1110/ps.44901. PMC 2374007
 . PMID 11369854.
. PMID 11369854. - ↑ Rao J, Cruz L (March 2013). "Effects of Confinement on the Structure and Dynamics of an Intrinsically Disordered Peptide: A Molecular-Dynamics Study". J. Of Phys. Chem B. 117 (14): 3707–3719. doi:10.1021/jp310623x.
- ↑ Uversky, Vladimir N (2013). "Digested disorder: Quarterly intrinsic disorder digest (January/February/March, 2013)". Intrinsically Disordered Proteins. 1: e25496. doi:10.4161/idp.25496.
- ↑ Susan Costantini; Ankush Sharma; Raffaele Raucci; Maria Costantini; Ida Autiero; Giovanni Colonna (March 2013). "Genealogy of an ancient protein family: the Sirtuins, a family of disordered members". BMC Evolutionary Biology. 13: 60. doi:10.1186/1471-2148-13-60. PMC 3599600
 . PMID 23497088.
. PMID 23497088.
External links
- Intrinsically disordered protein at Proteopedia
- MobiDB: a comprehensive database or intrinsic protein disorder annotations
- IDEAL - Intrinsically Disordered proteins with Extensive Annotations and Literature
- D2P2 Database of Disordered Protein Predictions
- Gallery of images of intrinsically disordered proteins
- First IDP journal covering all topics of IDP research
- IDP Journal
- Database of experimentally validated IDPs
- IDP ensemble database