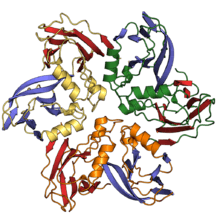
Jelly roll fold
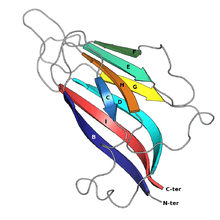
The jelly roll or Swiss roll fold is a protein fold or supersecondary structure composed of eight beta strands arranged in two four-stranded sheets. The name of the structure was introduced by Jane S. Richardson in 1981, reflecting its resemblance to the jelly or Swiss roll cake.[2] The fold is an elaboration on the Greek key motif and is sometimes considered a form of beta barrel. It is very common in viral proteins, particularly viral capsid proteins.[3][4] Taken together, the jelly roll and Greek key structures comprise around 30% of the all-beta proteins annotated in the Structural Classification of Proteins (SCOP) database.[5]
Structure
The basic jelly roll structure consists of eight beta strands arranged in two four-stranded antiparallel beta sheets which pack together across a hydrophobic interface. The strands are traditionally labeled B through H for the historical reason that the first solved structure, of a jelly roll capsid protein from the tomato bushy stunt virus, had an additional strand A outside the fold's common core.[6][7] The sheets are composed of strands BIDG and CHEF, folded such that strand B packs opposite strand C, I opposite H, etc.[4][8]
Viral proteins
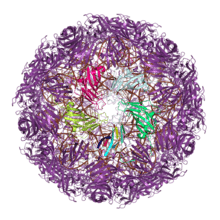
A large number of viruses build their exterior capsids from proteins containing either a single or a double jelly roll fold. This shared capsid architecture is thought to reflect ancient evolutionary relationships, possibly dating to before the last universal common ancestor (LUCA) of cellular life.[8][9][10] (Enclosed protein capsids themselves likely evolved at least twice, as other viral lineages use evolutionarily unrelated proteins to build their capsids.[9][11])
Single jelly roll capsid proteins
Single jelly roll capsid (JRC) proteins are found in at least sixteen distinct viral families, mostly with icosahedral capsid structures and including both RNA viruses and DNA viruses.[12] However, the majority of the single JRC viruses are positive-sense single-stranded RNA viruses, and the only double-stranded DNA viruses with single-JRC capsids are the Papillomaviridae and Polyomaviridae, both of which are quite small. The architecture of the assembled capsid orients the axis of the jelly roll parallel or "horizontally" relative to the capsid surface.[11]
Double jelly roll proteins
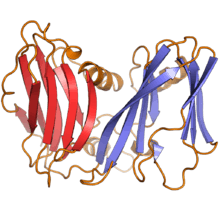
Double jelly roll capsid proteins consist of two single jelly roll folds connected by a short linker region. They are found exclusively in double-stranded DNA viruses of at least nine different viral families, including viruses that infect all domains of life, and spanning a large capsid size range.[4][11] In the double jelly roll capsid architecture, the jelly roll axis is oriented perpendicular or "vertically" relative to the capsid surface. Double jelly roll proteins are believed to have evolved from single jelly roll proteins by gene duplication; however, because of the major differences in architecture and assembly, it is unclear whether the double jelly roll capsid evolved directly from a single jelly roll capsid or if the two capsid forms represent distinct lineages of adaptation from common ancestral jelly roll proteins.[11][14] However, the degree of structural similarity among double-jelly-roll virus capsids has led to the conclusion that these viruses likely have a common evolutionary origin despite their diversity in size and in host range; this has become known as the PRD1-adenovirus lineage.[14] Although most members of this group are icosahedral, a few families such as the Poxviridae and Ascoviridae have oval or brick-shaped mature virions; poxviruses such as Vaccinia undergo dramatic conformational changes mediated by highly derived double jelly roll proteins during maturation and likely derive from an icosahedral ancestor.[11][15] Shared double-jelly-roll capsid proteins, along with other homologous proteins, have also been cited in support of the proposed order Megavirales containing the nucleocytoplasmic large DNA viruses (NCLDV).[16]
Double jelly roll proteins have not been observed in cellular proteins; they appear to be unique to viruses.[11] For this reason, detecting clear homology to double jelly roll proteins in the sequences of polinton/Maverick transposable elements widespread in eukaryotic genomes is considered evidence of these genetic elements' close evolutionary relationship to viruses.[17]
Non-capsid proteins
Single jelly rolls also occur in non-capsid viral proteins, including minor components of the assembled virion as well as non-virion proteins such as polyhedrin.[11]
Cellular proteins
While double jelly rolls are not found in proteins of cellular origin, single jelly rolls do occur.[11] One such class of cellular proteins is the nucleoplasmins, which serve as molecular chaperone proteins for histone assembly into nucleosomes. The N-terminal domain of nucleoplasmins possesses a single jelly roll fold and assembled into a pentamer.[18] Similar structures have since been reported in additional groups of chromatin remodeling proteins.[19] Jelly roll motifs with identical beta-sheet connectivity are also found in tumor necrosis factor ligands[20] and proteins from the bacterium Yersinia pseudotuberculosis that belong to a class of viral and bacterial proteins known as superantigens.[21][22]
More broadly, the members of the extremely diverse cupin superfamily are also often described as jelly rolls; though the common core of the cupin domain structure contains only six beta strands, many cupins have eight.[23] Examples include the non-heme dioxygenase enzymes[24][25] and JmjC-family histone demethylases.[26][27]
Evolution
Comparative studies of proteins classified as jelly roll and Greek key structures suggest that the Greek key proteins evolved significantly earlier than their more topologically complex jelly roll counterparts.[5] Structural bioinformatics studies comparing virus capsid jelly-roll proteins to other proteins of known structure indicates that the capsid proteins form a well-separated cluster, suggesting that they are subject to a distinctive set of evolutionary constraints.[4] One of the most notable features of viral capsid jelly roll proteins is their ability to form oligomers in a repeated tiling pattern to produce a closed protein shell; interestingly, the cellular proteins that are most similar in fold and topology are mostly also oligomers.[4]
History and nomenclature
The name "jelly roll" was first used for the structure composed of an elaboration on the Greek key motif by Jane S. Richardson in 1981 and was intended to reflect the structure's resemblance to a jelly or Swiss roll cake.[2] The structure has been given a variety of descriptive names, including a wedge, beta barrel, and beta roll. The edges of the two sheets do not meet to form regular hydrogen bonding patterns, and so it is often not considered to be a true beta barrel.[3] Cellular proteins containing jelly roll-like structures may be described as a cupin fold, a JmjC fold, or a double-stranded beta helix.[25]
External links
- Antiparallel β Domains, a section from Anatomy and Taxonomy of Protein Structure by Jane S. Richardson
- The Jelly Roll of Life by Jacqueline Humphries at Small Things Considered, a blog sponsored by the American Society for Microbiology
References
- 1 2 Larson, Steven B.; Day, John S.; McPherson, Alexander (29 August 2014). "Satellite tobacco mosaic virus refined to 1.4 Å resolution". Acta Crystallographica Section D. 70 (9): 2316–2330. doi:10.1107/S1399004714013789.
- 1 2 Richardson, JS (1981). "The anatomy and taxonomy of protein structure". Advances in Protein Chemistry. Advances in Protein Chemistry. 34: 167–339. doi:10.1016/S0065-3233(08)60520-3. ISBN 9780120342341. PMID 7020376.
- 1 2 Chelvanayagam, Gareth; Heringa, Jaap; Argos, Patrick (November 1992). "Anatomy and evolution of proteins displaying the viral capsid jellyroll topology". Journal of Molecular Biology. 228 (1): 220–242. doi:10.1016/0022-2836(92)90502-B.
- 1 2 3 4 5 Cheng, Shanshan; Brooks, Charles L.; Livesay, Dennis R. (7 February 2013). "Viral Capsid Proteins Are Segregated in Structural Fold Space". PLoS Computational Biology. 9 (2): e1002905. Bibcode:2013PLSCB...9E2905C. doi:10.1371/journal.pcbi.1002905. PMID 23408879.
- 1 2 Edwards, Hannah; Abeln, Sanne; Deane, Charlotte M.; Orengo, Christine A. (14 November 2013). "Exploring Fold Space Preferences of New-born and Ancient Protein Superfamilies". PLoS Computational Biology. 9 (11): e1003325. doi:10.1371/journal.pcbi.1003325. PMID 24244135.
- ↑ Harrison, S. C.; Olson, A. J.; Schutt, C. E.; Winkler, F. K.; Bricogne, G. (23 November 1978). "Tomato bushy stunt virus at 2.9 Å resolution". Nature. 276 (5686): 368–373. Bibcode:1978Natur.276..368H. doi:10.1038/276368a0.
- ↑ Rossmann, Michael G.; Abad-Zapatero, Celerino; Murthy, Mathur R.N.; Liljas, Lars; Jones, T. Alwyn; Strandberg, Bror (April 1983). "Structural comparisons of some small spherical plant viruses". Journal of Molecular Biology. 165 (4): 711–736. doi:10.1016/S0022-2836(83)80276-9.
- 1 2 Benson, Stacy D.; Bamford, Jaana K.H.; Bamford, Dennis H.; Burnett, Roger M. (December 2004). "Does Common Architecture Reveal a Viral Lineage Spanning All Three Domains of Life?". Molecular Cell. 16 (5): 673–685. doi:10.1016/j.molcel.2004.11.016.
- 1 2 Forterre, Patrick; Prangishvili, David (September 2009). "The origin of viruses". Research in Microbiology. 160 (7): 466–472. doi:10.1016/j.resmic.2009.07.008.
- ↑ Holmes, E. C. (30 March 2011). "What Does Virus Evolution Tell Us about Virus Origins?". Journal of Virology. 85 (11): 5247–5251. doi:10.1128/JVI.02203-10.
- 1 2 3 4 5 6 7 8 Krupovic, Mart; Bamford, Dennis H (August 2011). "Double-stranded DNA viruses: 20 families and only five different architectural principles for virion assembly". Current Opinion in Virology. 1 (2): 118–124. doi:10.1016/j.coviro.2011.06.001.
- ↑ Krupovic M (2013). "Networks of evolutionary interactions underlying the polyphyletic origin of ssDNA viruses". Current Opinion in Virology. 3 (5): 578–586. doi:10.1016/j.coviro.2013.06.010. PMID 23850154.
- 1 2 Abrescia, Nicola G.A.; Grimes, Jonathan M.; Kivelä, Hanna M.; Assenberg, Rene; Sutton, Geoff C.; Butcher, Sarah J.; Bamford, Jaana K.H.; Bamford, Dennis H.; Stuart, David I. (September 2008). "Insights into Virus Evolution and Membrane Biogenesis from the Structure of the Marine Lipid-Containing Bacteriophage PM2". Molecular Cell. 31 (5): 749–761. doi:10.1016/j.molcel.2008.06.026.
- 1 2 Krupovič, Mart; Bamford, Dennis H. (December 2008). "Virus evolution: how far does the double β-barrel viral lineage extend?". Nature Reviews Microbiology. 6 (12): 941–948. doi:10.1038/nrmicro2033.
- ↑ Bahar, Mohammad W.; Graham, Stephen C.; Stuart, David I.; Grimes, Jonathan M. (July 2011). "Insights into the Evolution of a Complex Virus from the Crystal Structure of Vaccinia Virus D13". Structure. 19 (7): 1011–1020. doi:10.1016/j.str.2011.03.023.
- ↑ Colson, Philippe; De Lamballerie, Xavier; Yutin, Natalya; Asgari, Sassan; Bigot, Yves; Bideshi, Dennis K.; Cheng, Xiao-Wen; Federici, Brian A.; Van Etten, James L.; Koonin, Eugene V.; La Scola, Bernard; Raoult, Didier (29 June 2013). ""Megavirales", a proposed new order for eukaryotic nucleocytoplasmic large DNA viruses". Archives of Virology. 158 (12): 2517–2521. doi:10.1007/s00705-013-1768-6.
- ↑ Krupovic, Mart; Bamford, Dennis H; Koonin, Eugene V (2014). "Conservation of major and minor jelly-roll capsid proteins in Polinton (Maverick) transposons suggests that they are bona fide viruses". Biology Direct. 9 (1): 6. doi:10.1186/1745-6150-9-6.
- ↑ Dutta, Shuchismita; Akey, Ildikó V.; Dingwall, Colin; Hartman, Kari L.; Laue, Tom; Nolte, Robert T.; Head, James F.; Akey, Christopher W. (October 2001). "The Crystal Structure of Nucleoplasmin-Core". Molecular Cell. 8 (4): 841–853. doi:10.1016/S1097-2765(01)00354-9. PMID 11684019.
- ↑ Edlich-Muth, Christian; Artero, Jean-Baptiste; Callow, Phil; Przewloka, Marcin R.; Watson, Aleksandra A.; Zhang, Wei; Glover, David M.; Debski, Janusz; Dadlez, Michal; Round, Adam R.; Forsyth, V. Trevor; Laue, Ernest D. (May 2015). "The Pentameric Nucleoplasmin Fold Is Present in Drosophila FKBP39 and a Large Number of Chromatin-Related Proteins". Journal of Molecular Biology. 427 (10): 1949–1963. doi:10.1016/j.jmb.2015.03.010.
- ↑ Bodmer, Jean-Luc; Schneider, Pascal; Tschopp, Jürg (January 2002). "The molecular architecture of the TNF superfamily". Trends in Biochemical Sciences. 27 (1): 19–26. doi:10.1016/S0968-0004(01)01995-8. PMID 11796220.
- ↑ Donadini, Roberta; Liew, Chu Wai; Kwan, Ann H.Y.; Mackay, Joel P.; Fields, Barry A. (March 2004). "Crystal and Solution Structures of a Superantigen from Yersinia pseudotuberculosis Reveal a Jelly-Roll Fold". Structure. 12 (1): 145–156. doi:10.1016/j.str.2003.12.002.
- ↑ Fraser, John D.; Proft, Thomas (October 2008). "The bacterial superantigen and superantigen-like proteins". Immunological Reviews. 225 (1): 226–243. doi:10.1111/j.1600-065X.2008.00681.x.
- ↑ Khuri, S; Bakker, FT; Dunwell, JM (April 2001). "Phylogeny, function, and evolution of the cupins, a structurally conserved, functionally diverse superfamily of proteins.". Molecular Biology and Evolution. 18 (4): 593–605. PMID 11264412.
- ↑ Ozer, Abdullah; Bruick, Richard K (March 2007). "Non-heme dioxygenases: cellular sensors and regulators jelly rolled into one?". Nature Chemical Biology. 3 (3): 144–153. doi:10.1038/nchembio863.
- 1 2 Aik, WeiShen; McDonough, Michael A; Thalhammer, Armin; Chowdhury, Rasheduzzaman; Schofield, Christopher J (December 2012). "Role of the jelly-roll fold in substrate binding by 2-oxoglutarate oxygenases". Current Opinion in Structural Biology. 22 (6): 691–700. doi:10.1016/j.sbi.2012.10.001. PMID 23142576.
- ↑ Chen, Zhongzhou; Zang, Jianye; Whetstine, Johnathan; Hong, Xia; Davrazou, Foteini; Kutateladze, Tatiana G.; Simpson, Michael; Mao, Qilong; Pan, Cheol-Ho; Dai, Shaodong; Hagman, James; Hansen, Kirk; Shi, Yang; Zhang, Gongyi (May 2006). "Structural Insights into Histone Demethylation by JMJD2 Family Members". Cell. 125 (4): 691–702. doi:10.1016/j.cell.2006.04.024. PMID 16677698.
- ↑ Klose, Robert J.; Zhang, Yi (7 March 2007). "Regulation of histone methylation by demethylimination and demethylation". Nature Reviews Molecular Cell Biology. 8 (4): 307–318. doi:10.1038/nrm2143.