Regulatory enzyme
A regulatory enzyme is an enzyme in a biochemical pathway which, through its responses to the presence of certain other biomolecules, regulates the pathway activity. This is usually done for pathways whose products may be needed in different amounts at different times, such as hormone production. Regulatory enzymes exist at high concentrations (low Vmax) so their activity can be increased or decreased with changes in substrate concentrations.
The enzymes which catalyse chemical reactions again and again are called regulatory enzymes.
Overview
Generally, it is considered that a hyperbolic structured protein in specific media conditions is ready to do its task, it is active, but some spe deactivation, are responsible for the regulation of some metabolism pathways. Regulatory enzymes are commonly the first enzyme in a multienzyme system: the product of the reaction catalyzed by the first enzyme is the substrate of the second enzyme, so the cell can control the amount of resulting product by regulating the activity of the first enzyme of the pathway.
There are many strategies of activation and deactivation of regulatory enzymes. Regulatory cific enzymes require an extra activation process and need to pass through some modifications in their 3D in order to become functional, for instance, catalyzing enzymes (regulatory enzymes). The regulation of the activation of these catalyzing enzymes is needed in order to regulate the whole reaction speed, so that it is possible to obtain the amount of product required at any time, that makes regulatory enzymes have a biological importance. Therefore, regulatory enzymes, by its controlled activation and are of two types: allosteric enzymes and covalently modulated enzymes; however, an enzyme can combine both types of regulation.
Allosteric enzymes
This type of enzymes presents two binding sites: the substrate of the enzyme and the effectors. Effectors are small molecules which modulate the enzyme activity; they function through reversible, non-covalent binding of a regulatory metabolite in the allosteric site (which is not the active site). When bound, these metabolites do not participate in catalysis directly, but they are still essential: they lead to conformational changes in a concrete part of the enzyme. These changes affect the overall conformation of the active site, causing modifications on the activity of the reaction.[1]
Properties
Allosteric enzymes are generally larger in mass than other enzymes. Different from having a single subunit enzyme, in this case they are composed of multiple subunits, which contain active sites and regulatory molecule binding sites.
They present a special kinetics: the cooperation. In here, configuration changes in each chain of the protein strengthen changes in the other chains. These changes occur at the tertiary and quaternary levels of organisation.
Based on modulation, they can be classified in two different groups:
- Homotropic allosteric enzymes: substrate and effector play a part in the modulation of the enzyme, which affects the enzyme catalytic activity.
- Heterotropic allosteric enzymes: only the effector performs the role of modulation.
Feedback inhibition
In some multienzyme systems, the enzyme is inhibited by the end product whenever its concentration is above the requirements of the cell. So, the velocity of the reaction can be controlled by the amount of product that is needed by the cell (the lower the requirement is, the slower the reaction goes).
Feedback inhibition is one of the most important function of proteins. Due to feedback inhibition, a cell is able to know whether the amount of a product is enough for its subsistence or there is a lack of the product (or there is too much product). The cell is able to react to this kind of situation in a mechanical way and solve the problem of the amount of a product. An example of feedback inhibition in human cells is the protein aconitase (an enzyme that catalyses the isomeration of citrate to isocitrate). When the cell needs iron, this enzyme loses the iron molecule and its form changes. When this happens, the aconitase is converted to IRPF1, a translation repressor or mRNA stabilizer that represses the formation of iron-binding proteins and favours formation of proteins that can get iron from the cell's reservations [1][2]
Covalently modulated enzymes
Here, the active and inactive form of the enzyme are altered due to covalent modification of their structures which is catalysed by other enzymes. This type of regulation consists of the addition or elimination of some molecules which can be attached to the enzyme protein. The most important groups that work as modifiers are phosphate, methyl, uridine, adenine and adenosine diphosphate ribosyl. These groups are joined to or eliminated from the protein by other enzymes. The most remarkable covalent modification is phosphorylation.
Phosphorylation
Phosphorylation is the addition of phosphate groups to proteins, which is the most frequent regulatory modification mechanism in our cells. This process takes place in prokaryotic and eukaryotic cells (in this type of cells, a third or a half of the proteins experience phosphorylation). Because of its frequency, phosphorylation has a lot of importance in regulatory pathways in cells.
The addition of a phosphoryl group to an enzyme is catalysed by kinase enzymes, while the elimination of this group is catalysed by phosphatase enzymes. The frequency of phosphorylation as a regulatory mechanism is due to the ease of changing from phosphorylated form to dephosphorylated form.
Phosphorylation or dephosphorylation make the enzyme be functional at the time when the cell needs the reaction to happen. The effects produced by the addition of phosphoryl groups that regulate the kinetics of a reaction can be divided in two groups:
- Phosphorylation changes the conformation of an enzyme to a more active or inactive way (e.g. regulation of glycogen phosphorylase). Each phosphate group contains two negative charges, so the addition of this group can cause an important change in the conformation of the enzyme. The phosphate can attract positively charged amino acids or create repulsive interactions with negatively charged amino acids. These interactions can change the conformation and the function of the enzyme. When a phosphatase enzyme removes the phosphate groups, this enzyme returns to its initial conformation.
- Phosphorylation modifies the affinity of the enzyme to the substrate (e.g. phosphorylation of isocitrate dehydrogenase creates electrostatic repulsion which inhibits the union of the substrate to the active center). Phosphorylation can take place in the active center of the enzyme. It can change the conformation of this active center, so it can recognize the substrate or not. Also, the ionized phosphate can attract some parts of the substrate, which can join to the enzyme.
Phosphorylation and dephosphorylation may take place as a result of the response to signals that warn about a change in the cell state. This means that some pathways where regulatory enzymes participate are regulated by phosphorylation after a specific signal: a change in the cell.
Some enzymes can be phosphorylated in multiple sites. The presence of a phosphoryl group in a part of a protein may depend on the folding of the enzyme (which can make the protein more or less accessible to kinase proteins) and the proximity of other phosphoryl groups.[1] [3][4]
Proteolysis
.jpg)
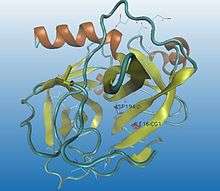
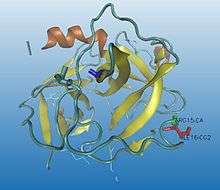
Some enzymes need to go through a maturation process to be activated. A precursor (inactive state, better known as zymogen) is first synthesized, and then, by cutting some specific peptide bonds (enzymatic catalysis by hydrolytic selective split), its 3D conformation is highly modified into a catalytic functional status, obtaining the active enzyme.
Proteolysis is irreversible and normally a non-specific process. The same activator can modulate different regulatory enzymes : once trypsin is activated, it activates many other hydrolytic enzymes. Proteolysis can also be fast and simple so the hydrolysis of a single peptide bond can be enough to change the conformation of the protein and build an active zone, allowing the interaction between the enzyme and the substrate, for instance, chymotrypsin activation (as it can be seen in the images).
Many different types of proteins with different roles in metabolism are activated by proteolysis for big reasons:
- Powerful hydrolytic enzymes, for instance, digestive enzymes, are activated by proteolysis so we can ensure that they are unable to hydrolyze any unwilling protein until they get to the right place: hydrolyzing protein zymogens are synthesized at the pancreas and accumulated in vesicles where they remain harmless. When they are needed, some hormonal or nervous stimulus triggers the release of the zymogens right to the intestine and they are activated.
- Some eventual responses must be immediate so enzymes that catalyze those reactions need to be prepared but not active, for that reason a zymogene is synthesized and stays ready for being rapidly activated. Coagulation response is based on enzymatic cascade proteolysis maturation. So, by activating one first catalyzing enzyme a big amount of the following enzymes is activated and the amount of product required is achieved as it is needed.
- Connective tissues proteins as collagen (zymogen: procolagen), hormones like insulin (zymogen: proinsulin) and proteins involved in development processes and apoptosis (programmed cell death) are activated by proteolysis too.
Proteolysis is irreversible, which implies the need of a process of enzyme deactivation. Specific inhibitors, analogous to the substrate, will strongly join the enzyme, blocking the substrate to join the enzyme. This union may last for months.[1][5]
References
- 1 2 3 4 Nelson, DL; Cox, MM (2009). Lehninger: Principios de bioquímica (5th ed.). Barcelona: Omega. pp. 220–228. ISBN 978-84-282-1486-5.
- ↑ Copley, SD (July 2012). "Moonlighting is Mainstream: Paradigm Adjustment Required". BioEssays. 34 (7): 578–588. doi:10.1002/bies.201100191.
- ↑ Alberts, B; Johnson, A (2008). Molecular Biology of the Cell (5th ed.). New York: Garland Science (GS). pp. 175–176. ISBN 0-8153-4106-7.
- ↑ Murray, RK; Bender, DA; Botham, KM; Kennely, PJ; Rodwell, VW; Weil, PA (2010). Harper. Bioquímica ilustrada (28th ed.). Mexico DF: Mc Graw Hill. pp. 80–81. ISBN 978-0-07-162591-3.
- ↑ Stryer, L; Berg, JM; Tymoczko, JL (2012). Biochemistry (Seventh ed.). New York: Palgrave, Macmillan. pp. 312–324. ISBN 978-1-4292-7635-1.